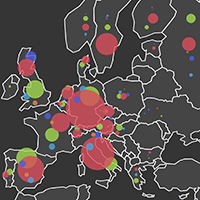
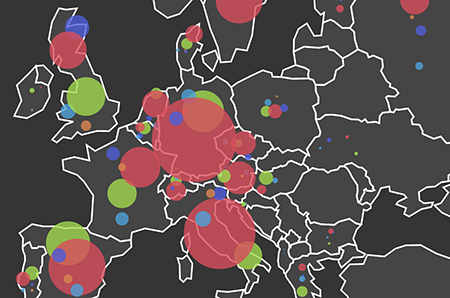
 This image describes the number and types of pathogens found in EU countries. The size of each circle is proportional to the number of pathogen species the EID2 team has evidence of, and the different coloured circles represent: bacteria (greenish yellow), fungi (red), helminths (purple blue), protozoa (orange), and viruses (blue).
This image describes the number and types of pathogens found in EU countries. The size of each circle is proportional to the number of pathogen species the EID2 team has evidence of, and the different coloured circles represent: bacteria (greenish yellow), fungi (red), helminths (purple blue), protozoa (orange), and viruses (blue).
Copyright: Dr Maya Wardeh, LUCINDA team
Researchers at the University of Liverpool’s Institute of Infection and Global Health are building the world’s most comprehensive database describing human and animal pathogens, which can be used to prevent and tackle disease outbreaks around the globe.
The Enhanced Infectious Diseases (EID2) database has been developed by the Liverpool University Climate and Infectious Diseases of Animals (LUCINDA) team and is funded by a BBSRC Strategic Tools and Resources Development Fund grant.
Huge benefits
Effectively mapping the relationships between human and animal diseases and their hosts, disease-causing pathogens and the ways in which pathogens are transmitted can offer huge benefits when it comes to knowing what the disease risks are in a population or geographical area, and how best to manage and eliminate them.
The EID2 team realised that there was a potential treasure trove of data already available in the scientific literature and in pre-existing databases, which was just waiting to be mined for useful insights – a ‘Big Data’ approach.
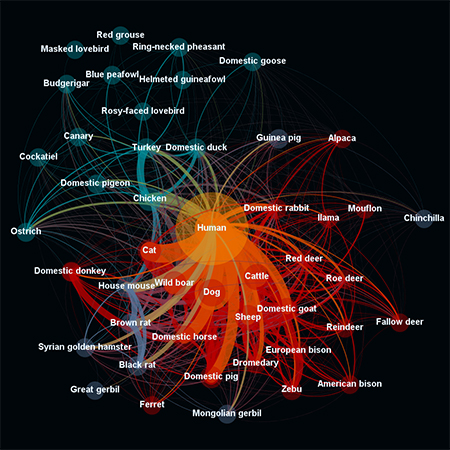 This depicts the relationship between information currently in the EID2 on domestic animal and human host species. The size of the nodes is relevant to the number of pathogen species found for each host; the arrows linking these nodes show the number of pathogen species shared by each pair of hosts (the thicker the arrow, the greater the number of pathogen species); and the colour of the nodes is related to the type of host (humans, other mammals, birds, rodents). Copyright: Dr Maya Wardeh, LUCINDA team
This depicts the relationship between information currently in the EID2 on domestic animal and human host species. The size of the nodes is relevant to the number of pathogen species found for each host; the arrows linking these nodes show the number of pathogen species shared by each pair of hosts (the thicker the arrow, the greater the number of pathogen species); and the colour of the nodes is related to the type of host (humans, other mammals, birds, rodents). Copyright: Dr Maya Wardeh, LUCINDA team
The emphasis on Big Data has increased recently because people have realised that the data that they have collected routinely, if used cleverly, can contain much more useful and potentially extra information than previously thought.
By using openly accessible information in a new way, data from EID2 has been used in work to trace the history of human and animal diseases, to predict the effects of climate change on pathogens, to produce maps of which diseases are most likely in some areas and to categorise the complex relationships between human and animal carriers and hosts of numerous pathogens.
Matchless in scale
Epidemiologist Dr Marie McIntyre, one of the EID2 team, said: “The database is matchless in scale, and has the capacity to hold data on all known human and animal pathogens, when detailed information becomes available.”
A longer version of this article was originally published on the Biotechnology and Biological Sciences Research Council (BBSRC) website.
Read the full version here.