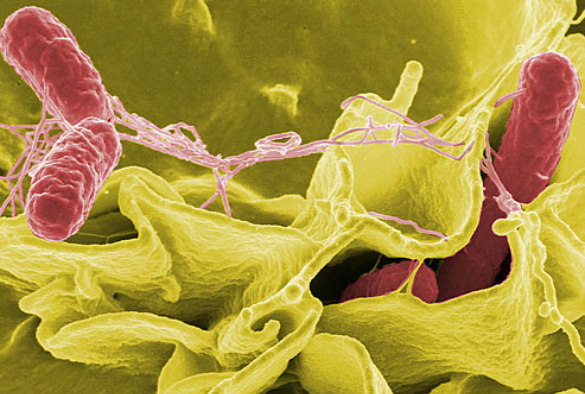
The first global-scale genetic study of Salmonella Enteritidis bacteria, which is a major cause of blood poisoning and death in Africa and food poisoning in the Western world, has discovered that there are in fact three separate types. Scientists at the Wellcome Trust Sanger Institute, Liverpool School of Tropical Medicine and University of Liverpool found two novel African types, which looked the same, but were genetically different from the Western type.
Reported in Nature Genetics, the study showed the African types had developed resistance to many antibiotics and behaved slightly differently to the type commonly found in the West. This suggests a need to understand where in the African environment the bacteria live, in order to prevent this disease.
Neglected Tropical Disease
Once Salmonella reaches the bloodstream it’s known as invasive non-typhoidal Salmonella (iNTS) disease, a serious and Neglected Tropical Disease. Last year, it was estimated to cause 680,000 per year deaths worldwide, more than half of which were in Africa.
The scientists sequenced 675 isolates of S. Enteritidis from 45 countries and six continents. Analysis of the Salmonella genomes revealed three major types – a common global one and two novel African types. Routine microbiological testing is not able to distinguish between the S. Enteritidis circulating in Africa and the rest of the world.
Identifying these new types is important because the common global type of S. Enteritidis, is normally associated with poultry and predominantly infects the intestine, causing diarrhoea. However, in Africa the two newly identified types are a major cause of blood poisoning and death, because in people with weak immune systems, Salmonellae are able to pass with greater ease from the gut into the bloodstream. The environmental reservoir from which these African bacteria are transmitted to people is unknown.
Antibiotic resistance
The study also showed that the two African types carried more of the genes that give them resistance to common antibiotics. These strains do not respond to the antibiotics commonly available, and have to be treated with cephalosporins or ciprofloxacin, antibiotics of last resort in many African settings.
Dr Nick Feasey, first author from the Liverpool School of Tropical Medicine and the Sanger Institute, said: “ iNTS disease is a huge problem in Africa, but it is difficult to identify due to lack of laboratory capacity and difficult to treat due antibiotic resistance. Using whole genome sequencing of S. Enteritidis bacteria from different settings we have revealed three distinct types of this bacteria, which was previously thought to have only one type. In the two novel types of Salmonella Enteritidis we found resistance genes to antibiotics such as amoxicillin and chloramphenicol, that are still widely used in Africa and it may be only a matter of time before resistance to the last line antibiotic, cephalosporin, spreads. We urgently need to find strategies to eradicate this disease.”
Professor Melita Gordon, joint lead author from the University of Liverpool’s Institute of Infection and Global Health, said: “To help prevent this disease, further investigations are urgently needed into where these African types reside in the environment and how they are passed on to or between people. It is possible that the Salmonella lives in standing water, so improving access to sanitation and clean water is extremely important. A human vaccine is also under development. Our research in Malawi, at the Malawi-Liverpool-Wellcome Trust (MLW) Clinical Research Programme, will hopefully yield and implement relevant public health strategies to prevent this disease.”
Professor Nick Thomson, joint lead author from the Sanger Institute, said: “This study highlights a very important issue; that a relatively mild version of a bacterium can evolve into a more dangerous pathogen under the right conditions. A combination of HIV, antibiotic resistance and lack of health care and sanitation has facilitated the emergence of Salmonella as a deadly disease in Africa and its importance was not originally recognised. The more we look, the more we understand the potential for bacteria which cause mild disease in rich settings to emerge as causes of highly deadly disease in Africa. Hence a critical first step in tackling diseases such as this is to be able to recognise the different ‘types’, which will in turn allow us to better understand how bacteria can exploit different ecological niches to which the global human population is exposed”.
The paper ‘Distinct Salmonella Enteritidis lineages associated with enterocolitis in high-income settings and invasive disease in low-income settings’ is published in Nature Genetics [DOI: http://dx.doi.org/10.1038/ng.3644].